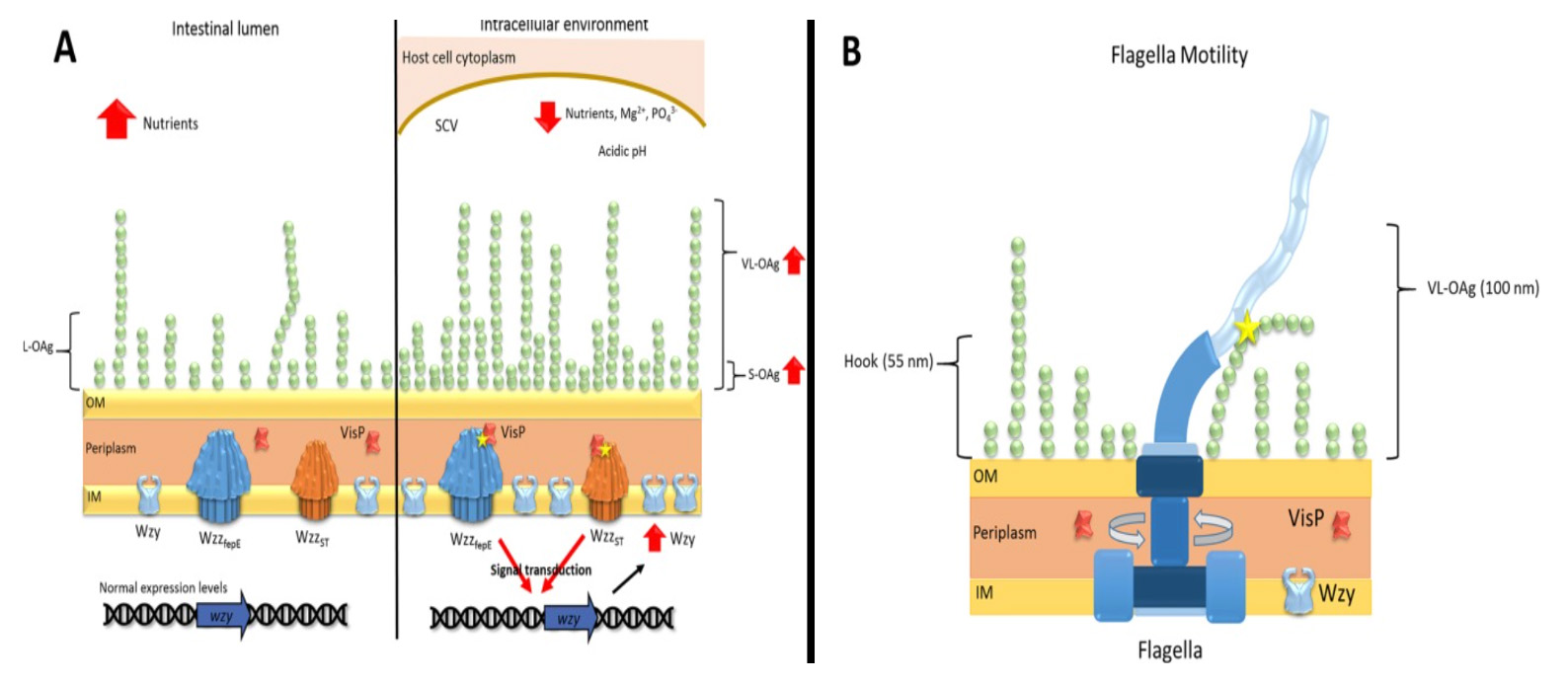
How do bacteria make us sick? A very simple and complicated question at the same time. Our research group has been successfully studying pathogenesis and chemical signaling in gram-negative bacteria over these last years in a diverse and integrated group of scientists from different levels and backgrounds. We investigate mechanisms of pathogenesis in distinct Salmonella and Escherichia coli pathogenic strains in the gut and urinary tract. Always exploring from basic to complex biological questions, how do they cause diseases, how do they adapt to hostile niches and evade the host response? Several bacterial pathogens rely on chemical signaling or quorum sensing to trigger their gene expression for distinct pathogenic pathways.
 |
|
The Basic Science Axis: Our main research projects cover distinct mechanisms such as |
Translational and Applied Science Axis: Development of alternative forms of treatment to circumvent |